
Center for Genome Innovation – Frequently Asked Questions
Main Location: 354 Mansfield Road (Beach Hall), Storrs Campus
Satellite Location: 400 Farmington Avenue, Farmington, UConn Health
Some common questions and answers are listed below. If you have a specific question or would like more information about any of the services provided by the CGI, contact Bo Reese.
When is the Center open?
Bo Reese and Lu Li have offices in the main laboratory (room 201) of the CGI, with either one or both of them present in the Center between 9:30am and 5:00pm. The Center is locked for security reasons during the times they are not there.
Can anyone use the instrumentation in the CGI?
With the appropriate training, anyone can use the instrumentation in the CGI. Depending on the instrumentation, the extent of training can range from a brief, one-on-one informal session to a three day workshop with other individuals. For example, training on the Agilent Bioanalyzer can be done in a one-on-one informal session, while usage of all NextGen sequencing instrumentation requires participation in a three day workshop or a one-on-one individual training. Even after training, users can request supervision if they are not comfortable working independently.
How do I reserve time to use the instruments in the CGI?
All equipment in the CGI is accompanied by a Google Calendar sign-up sheet. You can check the reservation schedule online by contacting either Bo Reese or Lu Li for access to the Google Calendars. Due to high usage volume of most of the equipment, we ask that individuals adhere to the schedules and cancel any reservations if they are no longer needed.
Are there fees associated with using any of the instrumentation?
Yes, for some instruments. The CGI requires usage access fees for some of the instruments located in the Center. The instruments included are:
- ABI 3130 Capillary Sequencer
- BioRad CFX 96 Real-Time System
- Roche 454 GS FLX+
- Illumina MiSeq
- Illumina NextSeq 500
- ABI SOLiD 5500xl
- Affymetrix Gene Chip
- Affymetrix Gene Atlas
- Fluidigm C1 Single-Cell Auto Prep, BioMark HD and Access Array systems
Instruments that do not require usage fees for access include:
- Agilent Bioanalyzer
- Agilent Tapestation
- BioRad Experion
- NanoDrop
- Hydroshear
In addition to instrument access fees, the CGI also offers fee-for-service for NextGen sequencing and genotyping services. A full list of services will be posted to the Fee-for-Services page when available, but in the meantime if you have questions or would like more information about any of the services provided by the CGI, contact Bo Reese.
How do I determine which Next Generation Sequencing platform is best for my experimental needs?
Most applications are supported by all NextGen sequencing platforms, like whole transcriptome RNA-Seq for example. Having your choice of which platform to use is great because each of the NextGen platforms has a unique way of presenting the data. Although there are certain experimental questions that heavily favor using one platform over another, there are others where the choice is not as clear. For many researchers, the choice in platform comes down to three main factors: genome target, desired genome coverage and cost. The information provided in the table below compares the most frequently used NextGen platforms based on these three (and other) factors.
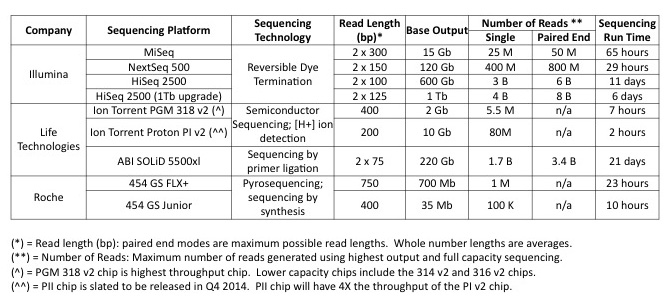
Because the details of each experiment are very different, we encourage you to reach out to us so that we can prepare a budget and sequencing coverage analysis for your experiment. Please contact Bo Reese with your questions.
Is there someone that can help me with NextGen Bioinformatic data analysis?
The CGI Bioinformatics Page provides links and references for some of the most commonly used bioinformatics tools on the web.
If you have a bioinformatics application in mind, but do not have the computing power to execute the application, the Biotechnology-Bioservices center may be able to help. Located in the Biology/Physics building on the Storrs campus, the Biotechnology-Bioservices center provides computational power and technical support to all faculty and students within the University system. They offer over 170 different bioinformatics applications accessible through a secure shell Unix command line. Please contact the Biotechnology-Bioservices center directly with your questions.